
22/03/2019
5th EU-ToxRisk General Assembly Meeting & 2nd Industry Stakeholder Meeting in February 2019
The european project EU-ToxRisk: "an integrated european 'flagship' program driving mechanism-based toxicity testing and risk assessment for the 21st century" is a H2020 supported collaborative project for the period 2016-2021 in which academia joins forces with small and medium-sized enterprises (SMEs), large industry, contract research organisations (CROs) and regulatory bodies to achieve a paradigm shift in toxicology towards a more efficient and animal-free chemical safety assessment. The GRIB takes part of this project as coordinator of WP4.
From 12-14 Feb. 2019, the EU-ToxRisk project hold its 5th General Assembly (GA) meeting, together with its 2nd Industry Stakeholders Meeting. This year's edition also took place in Egmond aan Zee, Netherlands. Altogether, some 120 people - including project members, advisory board members and external industry stakeholders - attended the event which was very successfull with interesting presentations, fruitful discussions, and instructive poster sessions.

05/03/2019
eTRANSAFE 5th Consortium Meeting in February 2019
The 5th eTRANSAFE Consortium Meeting held in Barcelona on the 20th and 21st of February 2019 to share the progress and discuss about achievements and next steps within the project.
The meeting started with a special workshop entitled "Preclinical and clinical terminology mapping" to set up strategies for terminology mapping for both preclinical and clinical studies, as well as identifying tools and resources. The second workshop of the day was focused on a review of the current version of workflows and how these will be taken up by the future knowledge hub to accommodate the needs of the end-users.
On the second day, two parallel workshops where held, i) Clinical data collection (trial data); and ii) Combining in vitro and gene expression data for elucidation of mechanisms of toxicity in vivo studies.
The meeting ended with the feedback from the Scientific Advisory Board members, who acknowledged the great involvement of the participants in the project and the eagerness to share expertise for the development of new strategies and the achievement of the project objectives.

04/02/2019
FAIRplus sets out to improve data sharing and reuse in life science research
The FAIRplus project has kicked-off to improve data management and data governance in European health research. The project aims to increase the discovery, accessibility and reusability of data from selected projects funded by the EU's Innovative Medicine Initiative (IMI), and internal data from pharmaceutical industry partners. It will also organise training for data scientists in academia, SMEs and pharmaceutical companies to enable wider adoption of best practises in life science data management.
Janssen Pharmaceutica NV is the pharmaceutical industry lead of the project with coordination provided by ELIXIR. The project involves 22 pharmaceutical, small company and academic partners from nine European countries. It is funded by the IMI as a public-private partnership and will run for three and a half years (2019-2022). European countries. It is funded by the IMI as a public-private partnership and will run for three and a half years (2019-2022). The Integrative Biomedical Informatics group of GRIB (IMIM - UPF) led by Laura I. Furlong and Ferran Sanz participates on this project and the main tasks foreseen are the interface with IMI projects in which we are involved (Open PHACTS, eTOX, eTRANSAFE, iPiE, EMIF), the involvement in the definition of the FAIRification processes of data in the following domains: preclinical toxicology, human safety, clinical data, drug impact in the environment, gene-disease associations and the organisation of events and other dissemination activities.
25/01/2019
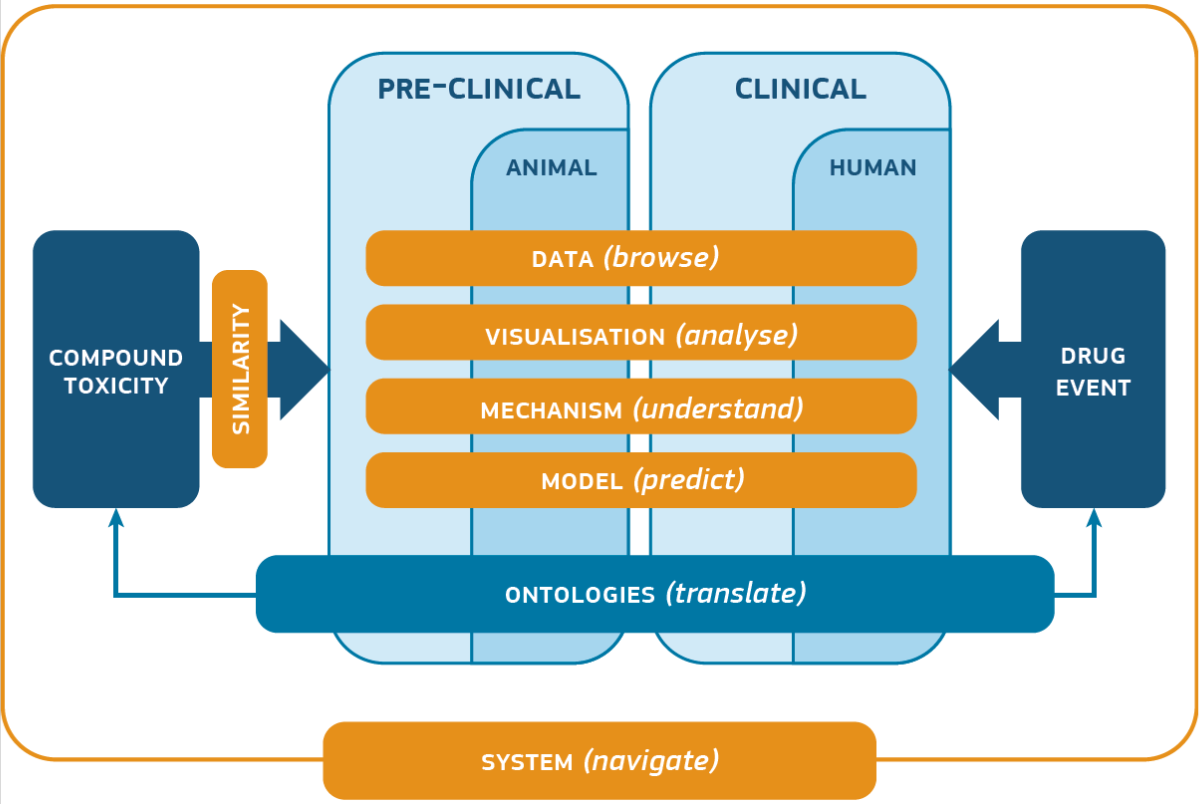
First year of the european project eTRANSAFE: read the keynote from Prof. Ferran Sanz, Project Coordinator
The eTRANSAFE project aims at enhancing the efficiency of translational safety assessment during the drug discovery and development process by means of the development of an integrative data infrastructure and innovative computational methods and tools. This infrastructure will be underpinned by legacy data sharing, the use of data standards, such as the SEND format for preclinical studies, and the implementation of a modular computational architecture.
eTRANSAFE started its journey in September 2017 and during the first year, we have set the foundations for the development of widely accepted guidelines on safe legacy data sharing, and we have been working in the extension of the preclinical database released in the previous IMI eTOX project, in terms of incorporation of individual animal data in SEND format together with structural and pharmacological information. In addition, identification of useful data available in the public domain, its integration with the data contributed from the EFPIA partners and the identification of potential challenges by means of prototypes and use cases have been the first steps to move forward the development of the eTRANSAFE Knowledge Hub.
The Project is pleased of the progress achieved and will continue advancing towards the final objective of supporting and improving the drug discovery and development.

16/01/2019
Ready, set, go for crossing the barrier!
Researchers at the CRG, GRIB and Helmholtz Center have discovered a new mechanism controlling the expression of a set of genes important for cell proliferation and tumour progression.
Genes contain all the information needed for the functioning of cells, tissues, and organs in our body. Gene expression, meaning when and how are the genes being read and executed, is thoroughly regulated like an assembly line with several things happening one after another.
Researchers at the Centre for Genomic Regulation (CRG) in collaboration with the Structural Bioinformatics group of GRIB (IMIM-UPF) led by Baldo Oliva and the Department of Molecular Epigenetics, Helmholtz Center Munich, Germany, have discovered a new step in this line, which controls the expression of some genes with an important role in cancer. "We observed that breast cancer cells need a particular modification to express a set of genes required for cellular proliferation and tumour progression," explains the CRG – Beatriu de Pinós postdoctoral researcher Priyanka Sharma, first author of the paper. "This modification allows the enzyme RNA polymerase II to overcome a pausing barrier and to continue to transcribe these genes," adds Sharma.
Reference article: Priyanka Sharma et al. "Arginine citrullination at the c-terminal domain controls RNA polymerase II transcription" Molecular Cell (2018) DOI: 10.1016/j.molcel.2018.10.01.
For press releases in spanish and catalan click here.

03/12/2018
Judith Pérez, winner of the collaborative poster session at the 2nd DCEXS PhD students' symposium
Congratulations to Judith Pérez, predoctoral researcher of the Integrative Biomedical Informatics Group of GRIB, awarded for the poster entitled "ResMarkerDB: a database of biomarkers of response to antibody therapy for breast and colorectal cancer". ResMarkerDB aims to promote the identification of existing and new actionable biomarkers of drug response in cancer and acts as a centralized repository that gathers information of biomarkers of response to therapeutic monoclonal antibodies for breast and colorectal cancer. Indeed, it integrates information from available public repositories and new data extracted and curated from the literature. Furthermore, ResMarkerDB allows exploration, visualization and prioritization of biomarker data and its response to therapy in a specific cancer type according to the evidence supporting their association and potential clinical usefulness. More information of this resource at http://resmarkerdb.org.
The poster session was part of the 2nd PhD students' symposium of the Department of Experimental and Health Sciences (DCEXS) held last 22&23rd of November in the PRBB. The DCEXS holds every year this symposium featuring leading topics in biomedical research, such as quantitative biology, genetics or neuroscience. Last year it was focused on doctoral students and this year it has been organized by a committee with representatives from all collectives.

28/11/2018
Optimizing collaboration between academia and the pharmaceutical industry to improve the toxicological evaluation of drugs
A new article by Manuel Pastor, Jordi Quintana and Ferran Sanz shares their experience in the eTOX project to improve the transfer of models from academia to industry.
Researchers Manuel Pastor, Jordi Quintana and Ferran Sanz have published an article in the journal Frontiers in Pharmacology in which they share their experience gained in the transfer of computational predictive models from academic to industrial environments. This experience was acquired in the already concluded eTOX project, funded by the Innovative Medicines Initiative (IMI). The main results of this project pave the way for a new model of collaboration by the pharmaceutical industries among themselves and also between them and the world of academia, by sharing data and knowledge in order to improve the toxicological evaluation of compounds for potential use in drugs.
"In this article we have shared our experience in the adaptation of computational tools for drug discovery generated in academia for use in the pharmaceutical industry. It is an aspect that is often overlooked in this type of projects", says Manuel Pastor, head of the GRIB's Pharmacoinformatics group. "We have described some of the problems identified and the solutions we apply to solve or mitigate them", he adds.
In the project, the pharmaceutical companies contributed data generated and stored internally to train predictive models. One of the problems encountered was the need to share confidential information, such as the structure of compounds under development. Given that such data are highly sensitive, the companies were opposed to letting this information out of their internal repositories.

21/11/2018
GRIB participated in the iPiE Forum Meeting held in Brussels from the 13th to the 15th of November
The iPiE consortium members met in Brussels, hosted by Janssen in its Diegem offices. The first day, the meeting was structured in workshops and general sessions on updates discussions. The second day counted with the attendance of the Scientific Advisory Board members of the project, to whom the last achievements and progress was presented. The last day was aimed to discuss on the prioritisation guidelines which are being developed in the final months of the project.
iPiE aims to develop predictive frameworks that use information from existing datasets on environmental fate and effects of Active Pharmaceutical Ingredients (APIs), toxicological studies, pharmacological mode of action and in silico models to support more intelligent environmental testing of pharmaceuticals. The iPiE project has delivered models to estimate concentrations of pharmaceuticals in surface waters, sediments and soils across Europe as well as the models for predicting effects of pharmaceuticals on aquatic organisms. One of the major iPiE achievements was the development and the release of the iPiEsys software which allows the database to be searched and predictive models to be run. The guidance on how the software system and associated predictive tools can be used is under development and it is expected to be delivered early next year.
More about the project at http://i-pie.org/

16/11/2018
Conocer la toxicidad para mejorar la seguridad
El éxito del proyecto eTOX, nacido en 2010 en el marco de la iniciativa europea IMI (Innovative Medicines Initiative) dio paso en 2017 a un nuevo proyecto, herencia del anterior, pero bastante más ambicioso: eTRANSAFE. En ambos casos, se trata de compartir datos de investigación relacionados con la toxicidad para mejorar la seguridad de nuevos compuestos en desarrollo aprovechando la información que las compañías farmacéuticas extraen de sus procesos.
Ferran Sanz, coordinador académico de ambos proyectos y director del GRIB, manifiesta que "estamos orgullosos de haber conseguido que las compañías accedieran a compartir sus datos preclínicos en el eTOX", aunque reconoce que "los dos primeros años costó un poco". Ahora, la segunda etapa va un paso más allá porque requiere poner en común datos de ensayos clínicos. No obstante, recuerda que sólo se comparten datos de toxicidad y seguridad; nunca de eficacia, que serían los más sensibles. La previsión es que la nueva base de datos sea mucho mayor, porque se hará con más agilidad gracias al SEND y porque amplía el espectro de datos a reunir: tanto preclínicos como clínicos.
Lee la entrevista completa publicada por Diario Médico en su último newsletter.

05/11/18
eTRANSAFE project at the IMI’s 10th Anniversary Celebration
eTRANSAFE was present at the IMI 10th Anniversary Scientific Symposium and the IMI Stakeholder Forum, both events were jointly held in Brussels in the framework of the IMI's 10th anniversary celebration. The IMI Stakeholder Forum held on the 24th of October had as theme 'The value of cross-sectoral health research and innovation'. eTRANSAFE Project Coordinator, Ferran Sanz , Director of GRIB (IMIM-UPF), participated in this event and took part in a session entitled "Case study: how to apply Technology convergence in safety", together with other members from Novartis, Mimetas, Sanofi and GIRP. During this session Prof. Sanz had the chance to introduce eTRANSAFE and to discuss on the benefits of data sharing contributing to the improvement of safety assessment.
IMI is funded jointly by the European Union (represented by the European Commission) and the European pharmaceutical industry (represented by EFPIA, the European Federation of Pharmaceutical Industries and Associations). The EU funding comes from Horizon 2020 and the EU's Seventh Framework Programme for Research (FP7). More info about IMI 10th Anniversary: here



